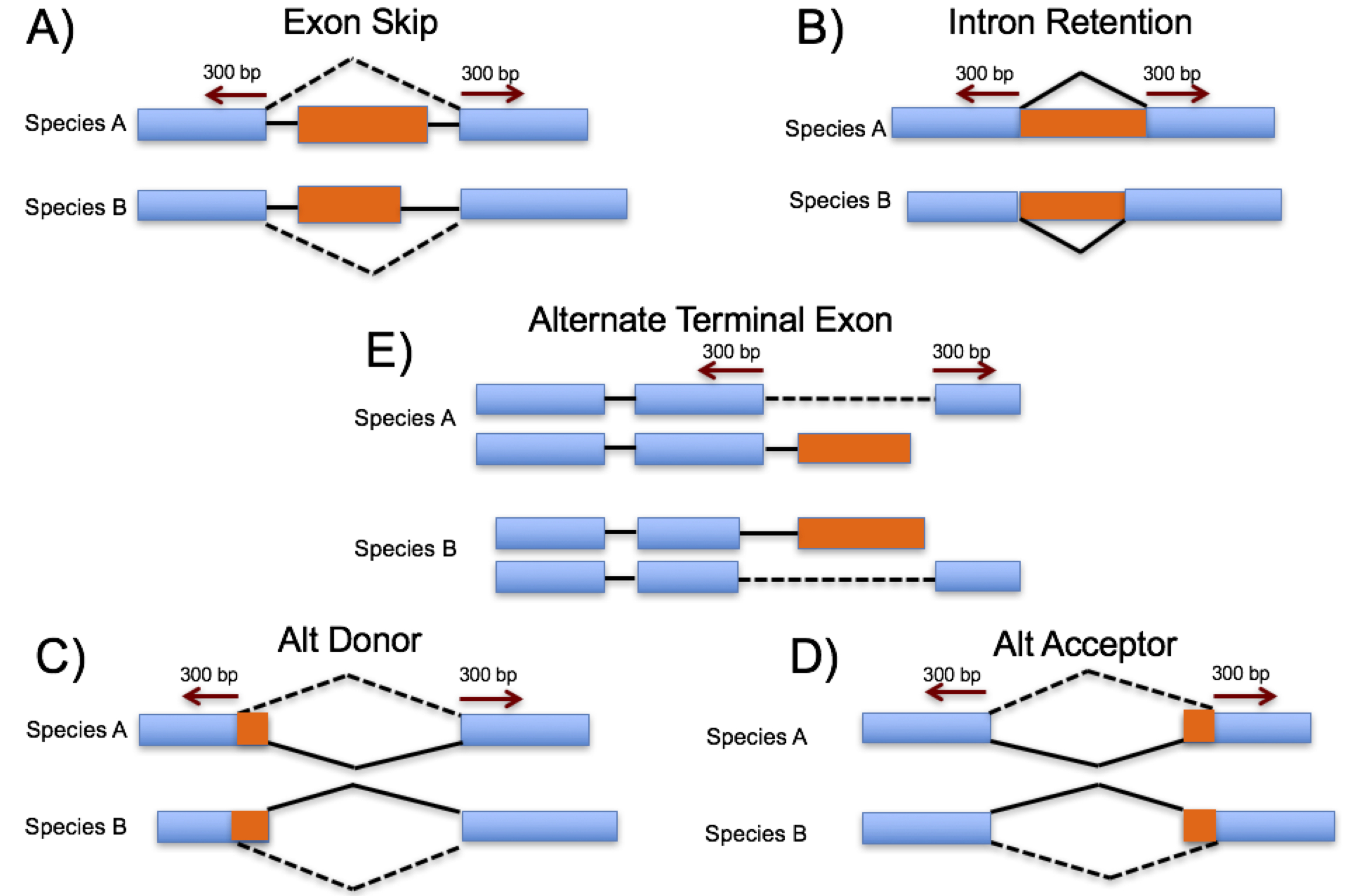
One difficulty when identifying and analyzing alternative splicing (AS) events in plants is distinguishing functional AS from splicing noise. One way to add confidence to the validity of a splice isoform is to observe that it is conserved across evolutionarily related species. We use a high throughput method to identify junction based conserved AS events from RNA-Seq data across nine plant species including: five grass monocots (maize, sorghum, rice, Brachpodium and foxtail millet), plus two non-grass monocots (bananan and African oil palm), the eudicot Arabidopsis and the basal angiosperm Amborella.
In total, 9,804 conserved AS events within 19,235 genes were identified conserved between 2 or more species studied. In grasses containing large regions of conserved synteny, the frequency of conserved AS events is twice that observed for genes outside of conserved synteny blocks.
In plant-specific RS and RS2Z subfamilies, we observe both conservation and divergence of AS events after the whole genome duplication in maize. In addition, plant-specific RS and RS2Z subfamilies are highly connected with R2R3-MYB in splicing networks.
Furthermore, we discovered that the network based on genes harboring conserved AS events is enriched for phosphatases, kinases and ubiquitylation genes, which suggests that AS may participate in regulating signaling pathways. These data lay the foundation for identifying and studying conserved AS events in the monocots, particularly across grass species, and this conserved AS resource identifies an additional layer between genotype to phenotype that may impact future crop improvement efforts.